What exactly is the D816V mutation and why does it matter? To answer that, we need to understand the basic pathway by which a cell expresses a gene.
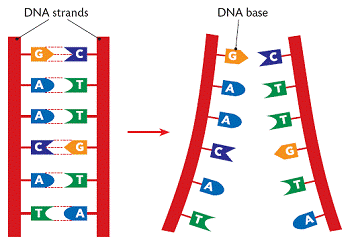
DNA (deoxyribonucleic acid) is the molecule that contains the genetic code for all known living organisms and some viruses. DNA is composed of two strands that wrap around each other in a double helix pattern. DNA is built out of nucleotides, molecules that contain energy. The nucleotides that build DNA are adenine (A), guanine (G), thymine (T) and cytosine (C). These nucleotides bond in specific pairs. This means that when one nucleotide in on one strand of DNA, there is a specific nucleotide on the other strand. A and T, and C and G specifically bond to each other. They are known as base pairs. DNA strands made up of base pairs are said to be “complementary.”
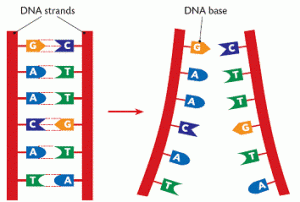
RNA (ribonucleic acid) is a more versatile nucleic acid that codes, regulates and expresses genes, amongst other things. It also has base pairs: adenine and uracil (U), and thymine and cytosine. These nucleotides can be complementary to DNA nucleotides. For example, an RNA adenine is complementary to a DNA thymine, and so on.
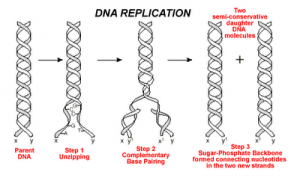
DNA replication is the process by which an exact copy of a piece of DNA is made. This happens when a cell divides. In replication, the DNA double helix “unzips,” or splits apart into two strands, the base pairs of which are not connected. Special enzymes move along each of the two split strands and place the appropriate nucleotides next to each strand to form base pairs. The end result of this is two double helices of DNA that are exact copies.
Some parts of DNA, called genes, tell the cell how to make proteins or RNA that has a specific function. (Sometimes RNA can also do this.) Genes tell the cell how to build and maintain the cell and allow it pass on traits to offspring. These proteins or RNA are made by expressing the gene. In gene expression, the information from the gene is turned into a “gene product,” that will be made into something useful for the cell.
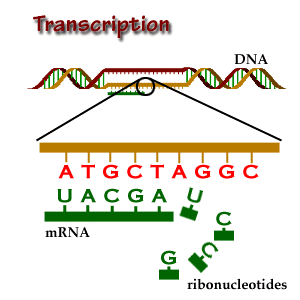
Transcription is the start of gene expression. Gene expression is very complicated and controlled by many mechanisms. Having a gene does not mean it will always be expressed. In transcription, a piece of DNA is copied into a complementary RNA strand. This RNA is called messenger RNA (mRNA.) This is a complicated process with several steps. Once a gene is translated, the mRNA with the gene code goes to the ribosome, a place in the cell that makes proteins. Proteins are made of amino acids.
So how exactly does the DNA code for the protein the ribosome will make? Let’s focus on that.
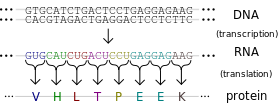
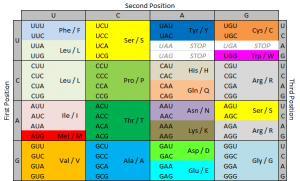
The ribosome reads the messenger RNA made from the DNA gene three nucleotides at a time. Again, when using the code to build a protein, the ribosome reads the code in blocks of three nucleotides. These blocks of three nucleotides are called “codons.” Every combination of three-nucleotides tells the ribosome to add a specific amino acid to the protein. The majority of genes are encoded using this same codon code. So by knowing the DNA sequence, we can anticipate the amino acids that build the protein encoded by the gene.
How does the ribosome know where to start? There’s a start codon. (And some other things also.)
How does the ribosome know where to stop? There’s a stop codon. (And some other things also.)
There are several types of genetic mutations, or alterations of the code from the one seen in most of the population. In a point mutation, a single nucleotide is changed. The D816V mutation is a point mutation.
We use a specific nomenclature to describe genetic mutations. Amino acids are often referred to with single letter codes for the sake of brevity. The amino acid aspartic acid is referred to as “D,” while the amino acid for valine is referred to as “V.” In the CKIT gene, the amino acid sequence Asp-Phe-Gly (aspartic acid – phenylalanine – glycine) is very important to the receptor being shaped the right way.
Aspartic acid is encoded by the RNA code “GAU” or “GAC.” In cells with the D816V mutation, this sequence is changed to “GUU” or “GUC.” The second base is changed from an A to a U. Doing this changes the amino acid encoded from aspartic acid (D) to valine (V). These amino acids are shaped differently, and because of this, the receptor is shaped differently and behaves differently. When the receptor is made with the amino acid aspartic acid in that place, SCF (stem cell factor) binds to the receptor and activates the cell, telling it not to die and to make more cells. When the receptor is made with the amino acid valine in that place, the receptor activates itself and SCF is not needed. It basically tells itself not to die and to make more cells repeatedly.